Predict from a Cross-Validated Joint Model Fit
predict.swjm_cv.RdComputes subject-specific predictions from a cross-validated swjm_cv
fit. The output differs by model:
Usage
# S3 method for class 'swjm_cv'
predict(object, newdata, times = NULL, ...)Arguments
- object
An object of class
"swjm_cv".- newdata
A numeric matrix or data frame of covariate values. Must have
pcolumns namedx1, ...,xp(if a data frame) or exactlypcolumns (if a matrix).- times
Numeric vector of evaluation times (JFM only). If
NULL, the observed event times from the training data are used.- ...
Currently unused.
Value
An object of class "swjm_pred", a list with:
- S_re
Matrix of readmission-free survival probabilities (rows = subjects, columns =
times).- S_de
Matrix of death-free survival probabilities.
- times
Numeric vector of evaluation times.
- lp_re
Linear predictors for readmission (\(\hat\alpha^\top z_i\)). For JSCM this is the log time-acceleration for the recurrence process.
- lp_de
Linear predictors for death (\(\hat\beta^\top z_i\)). For JSCM this is the log time-acceleration for the terminal process.
- time_accel_re
(JSCM only) \(e^{\hat\alpha^\top z_i}\): the multiplicative factor by which the recurrence time axis is scaled relative to baseline.
NULLfor JFM.- time_accel_de
(JSCM only) Analogous time-acceleration factor for the terminal process.
NULLfor JFM.- contrib_re
Matrix of per-predictor contributions \(\hat\alpha_j z_{ij}\) (rows = subjects, columns = covariates). For JFM these are log-hazard contributions; for JSCM they are log time-acceleration contributions.
- contrib_de
Analogous matrix for the terminal process.
Details
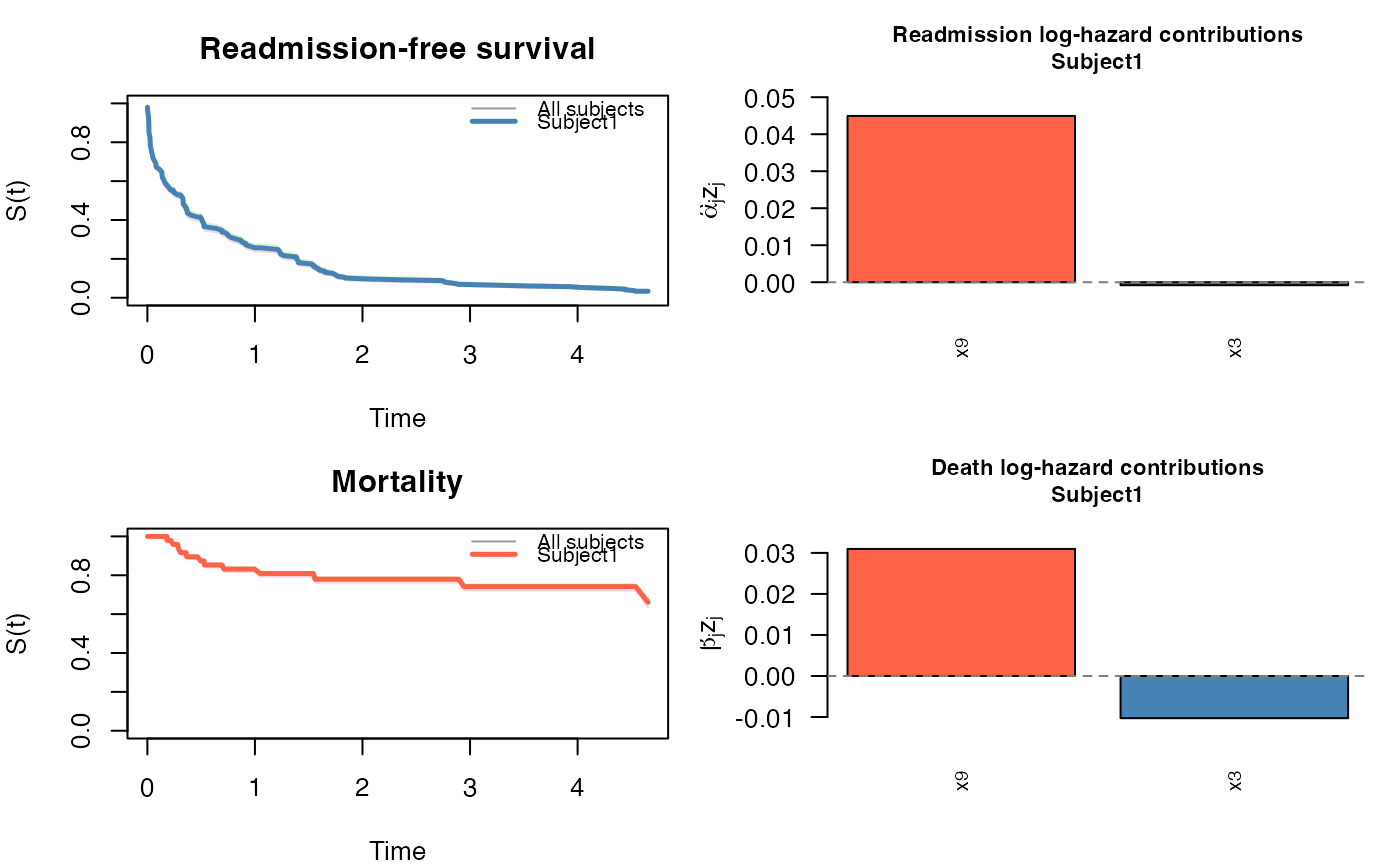
JFM — returns survival curves for both processes using the Breslow cumulative baseline hazards. For subject \(i\) with covariate vector \(z_i\): $$ S_{\text{re}}(t \mid z_i) = \exp\!\bigl(-\hat\Lambda_0^r(t)\, e^{\hat\alpha^\top z_i}\bigr), \quad S_{\text{de}}(t \mid z_i) = \exp\!\bigl(-\hat\Lambda_0^d(t)\, e^{\hat\beta^\top z_i}\bigr). $$
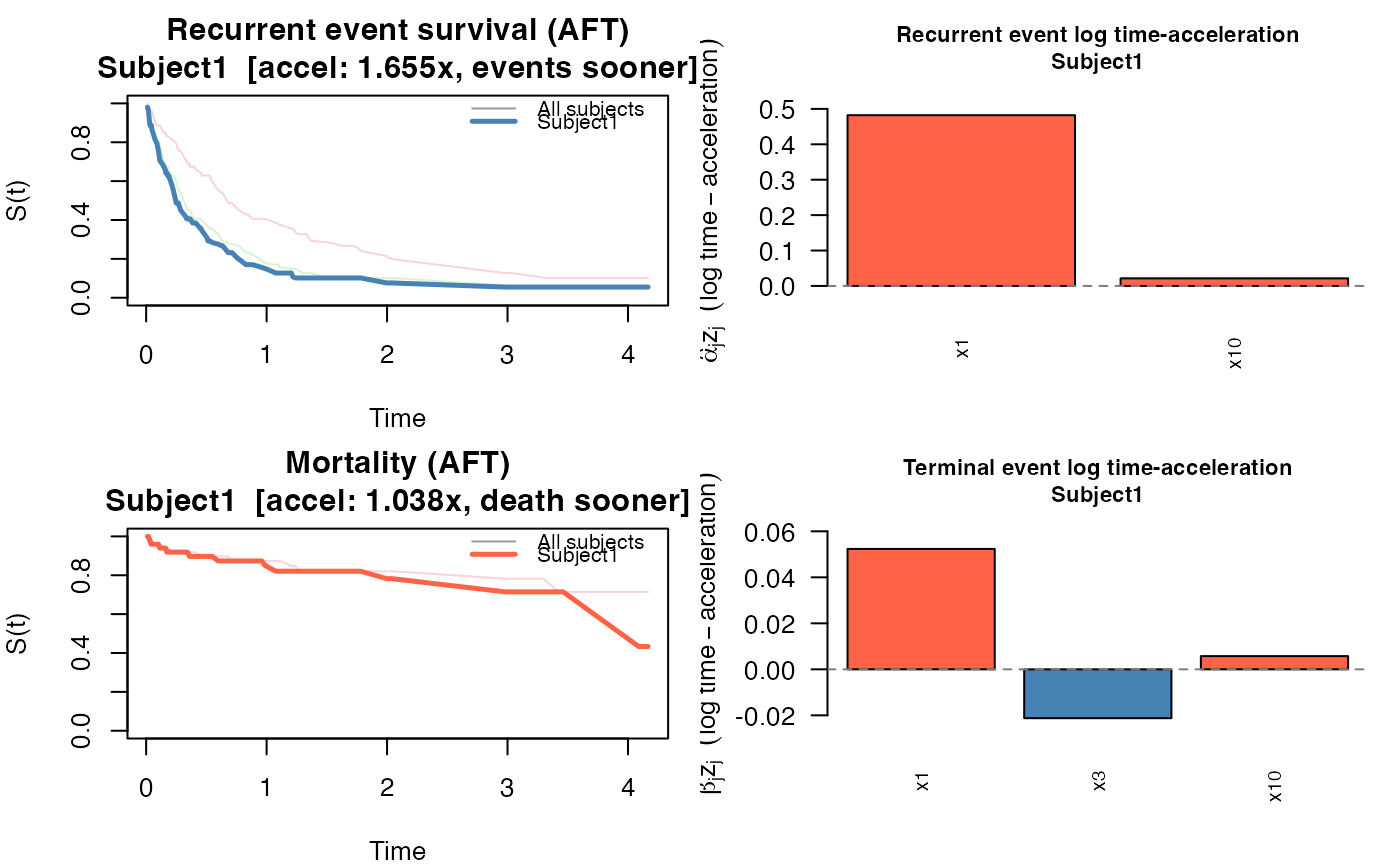
JSCM — returns survival curves for both processes using a Nelson-Aalen baseline on the accelerated time scale. For subject \(i\): $$ S_{\text{re}}(t \mid z_i) = \exp\!\bigl(-\hat\Lambda_0^r(t\, e^{\hat\alpha^\top z_i})\bigr), \quad S_{\text{de}}(t \mid z_i) = \exp\!\bigl(-\hat\Lambda_0^d(t\, e^{\hat\beta^\top z_i})\bigr). $$ The linear predictor \(\hat\alpha^\top z_i\) is a log time-acceleration factor: the recurrence process for subject \(i\) runs on a time axis scaled by \(e^{\hat\alpha^\top z_i}\) relative to the baseline. Each predictor contributes \(\hat\alpha_j z_{ij}\) to this log-scale factor, so \(e^{\hat\alpha_j z_{ij}}\) is the multiplicative factor on the time scale attributable to predictor \(j\) alone. Values greater than 1 accelerate events (shorter times); values less than 1 decelerate them (longer times).
Examples
# \donttest{
dat <- generate_data(n = 50, p = 10, scenario = 1, model = "jfm")
cv <- cv_stagewise(dat$data, model = "jfm", penalty = "coop",
max_iter = 100)
newz <- matrix(rnorm(30), nrow = 3, ncol = 10)
pred <- predict(cv, newdata = newz)
plot(pred)
 dat_jscm <- generate_data(n = 50, p = 10, scenario = 1, model = "jscm")
#> Call:
#> reReg::simGSC(n = n, summary = TRUE, para = para, xmat = X, censoring = C,
#> frailty = gamma, tau = 60)
#>
#> Summary:
#> Sample size: 50
#> Number of recurrent event observed: 81
#> Average number of recurrent event per subject: 1.62
#> Proportion of subjects with a terminal event: 0.22
#>
#>
cv_jscm <- cv_stagewise(dat_jscm$data, model = "jscm", penalty = "coop",
max_iter = 500)
pred_jscm <- predict(cv_jscm, newdata = newz)
plot(pred_jscm)
dat_jscm <- generate_data(n = 50, p = 10, scenario = 1, model = "jscm")
#> Call:
#> reReg::simGSC(n = n, summary = TRUE, para = para, xmat = X, censoring = C,
#> frailty = gamma, tau = 60)
#>
#> Summary:
#> Sample size: 50
#> Number of recurrent event observed: 81
#> Average number of recurrent event per subject: 1.62
#> Proportion of subjects with a terminal event: 0.22
#>
#>
cv_jscm <- cv_stagewise(dat_jscm$data, model = "jscm", penalty = "coop",
max_iter = 500)
pred_jscm <- predict(cv_jscm, newdata = newz)
plot(pred_jscm)
 # }
# }